CDORemap(): irregular regridding has problems when input dim number < 4
Hi @nperez
I found a problem following s2dverification#259 (closed). The data (the 1st data) in that issue work well with the irregular regridding, but not for data (the 2nd data) under /esarchive/exp/CMIP6/dcppA-hindcast/cmcc-cm2-sr5/cmip6-dcppA-hindcast_i1p1/DCPP/CMCC/CMCC-CM2-SR5/dcppA-hindcast/.
The problem only happens when the dimension number of input of CDORemap() is < 4, which also happened to the 1st data before (see the last discussion of s2dverification#259 (closed)). The main difference between 1st and 2nd data is i and j. The 1st data has j = 292; i = 362 while the 2nd data has i = 292; j = 362. It leads to the swapped latitude and longitude. I put the code below.
# 1st data (CORRECT)
path1 <- '/esarchive/exp/ecearth/a1ua/cmorfiles/DCPP/EC-Earth-Consortium/EC-Earth3/dcppA-hindcast/r1i1p1f1/Omon/$var$/gn/v20190713/$var$_*_s$sdate$-$member$_gn_$aux$.nc'
data1 <- Start(dataset = path1,
var = 'tos',
sdate = paste0(1960),
aux = indices(1),
aux_depends = 'sdate',
j = 'all',
i = 'all',
time = indices(1),
member = 'r1i1p1f1',
return_vars = list(j = NULL, i = NULL,
latitude = NULL, longitude = NULL),
retrieve = T)
lons1 <- attributes(data1)$Variables$common$longitude
lats1 <- attributes(data1)$Variables$common$latitude
dim(lons1)
dim(lats1)
dim(data1)
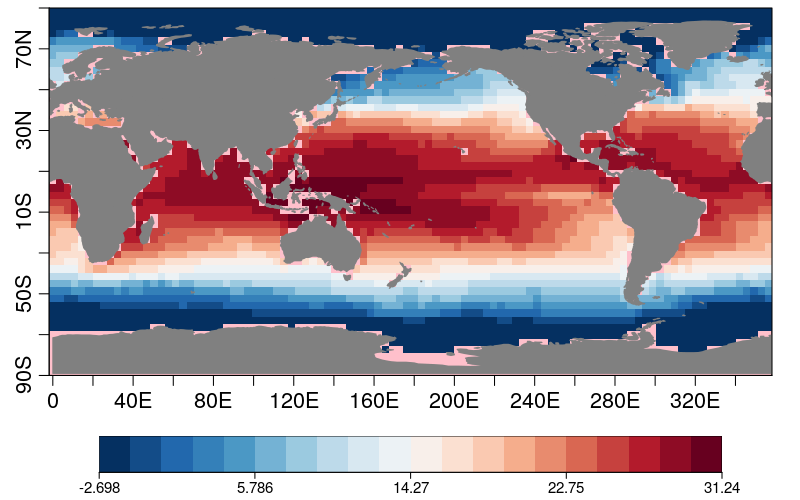
res1 <- CDORemap(drop(data1), lons1, lats1, grid = 'r100x50', method = 'bil', crop = FALSE)
dim(res1$data_array)
#lon lat
#100 50
s2dv::PlotEquiMap(t(res1$data_array), lon = res1$lons, lat = res1$lats)
# 2nd data (WRONG. If don't drop(data2), it will be correct.)
path2 <- paste0('/esarchive/exp/CMIP6/dcppA-hindcast/cmcc-cm2-sr5/cmip6-dcppA-hindcast_i1p1/',
'DCPP/CMCC/CMCC-CM2-SR5/dcppA-hindcast/$member$/Omon/$var$/gn/v20210312/',
'$var$_*_s$sdate$-$member$_gn_$aux$.nc')
data2 <- Start(dataset = path2,
var = 'tos',
sdate = '1960',
aux = 'all',
aux_depends = 'sdate',
j = 'all',
i = 'all',
time = indices(1),
member = 'r1i1p1f1',
return_vars = list(j = NULL, i = NULL,
latitude = NULL, longitude = NULL),
retrieve = T)
lons2 <- attributes(data2)$Variables$common$longitude
lats2 <- attributes(data2)$Variables$common$latitude
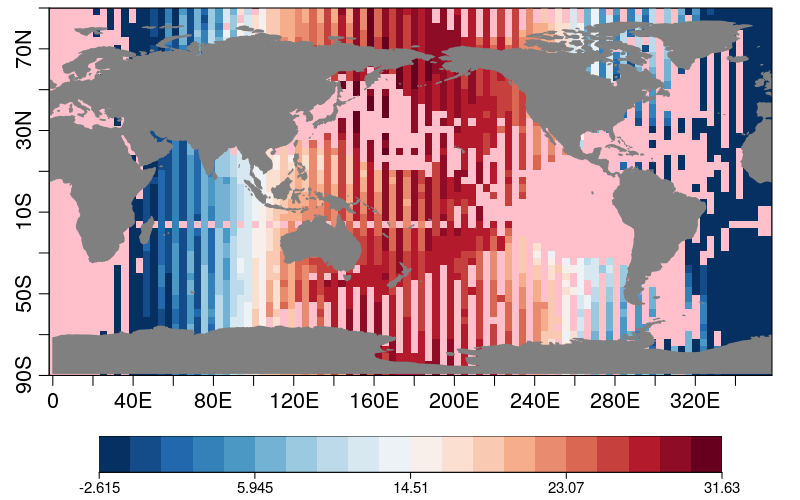
res2 <- s2dv::CDORemap(drop(data2), lons2, lats2, grid = 'r100x50', method = 'bil', crop = FALSE)
dim(res2$data_array)
#lat lon
# 50 100
s2dv::PlotEquiMap(res2$data_array, lon = res2$lons, lat = res2$lats)Cheers,
An-Chi