Interpolation of Decadal original files into a regular grids avoiding vertical empty lines and artifacts
Hi @rfernand
Finally, we are able to load and regrid IPSL files avoiding artifacts with the code:
library(startR)
library(s2dverification)
# Path to original cmorfiles:
path <- '/esarchive/exp/ipsl-cm6a-lr/cmip6-dcppA-hindcast_i1p1/original_files/cmorfiles/DCPP/IPSL/IPSL-CM6A-LR/dcppA-hindcast/r1i1p1f1/SImon/siconc/gn/v20200101/$var$_SImon_IPSL-CM6A-LR_dcppA-hindcast_s2016-r1i1p1f1_gn_201701-202612.nc'
# Regrid:
data <- Start(dat = path,
var = 'siconc',
x = indices(2:361),
y = indices(2:331),
time = 'all',
return_vars = list(nav_lat = NULL, nav_lon = NULL),
retrieve = TRUE)
nav_lon <- attributes(data)$Variables$common$nav_lon[2:361, 2:331]
nav_lat <- attributes(data)$Variables$common$nav_lat[2:361, 2:331]
res <- CDORemap(data[1,1, , ,5],
nav_lon,
nav_lat,
grid = 'r360x180',
method = 'bicubic', crop = FALSE)
dev.new()
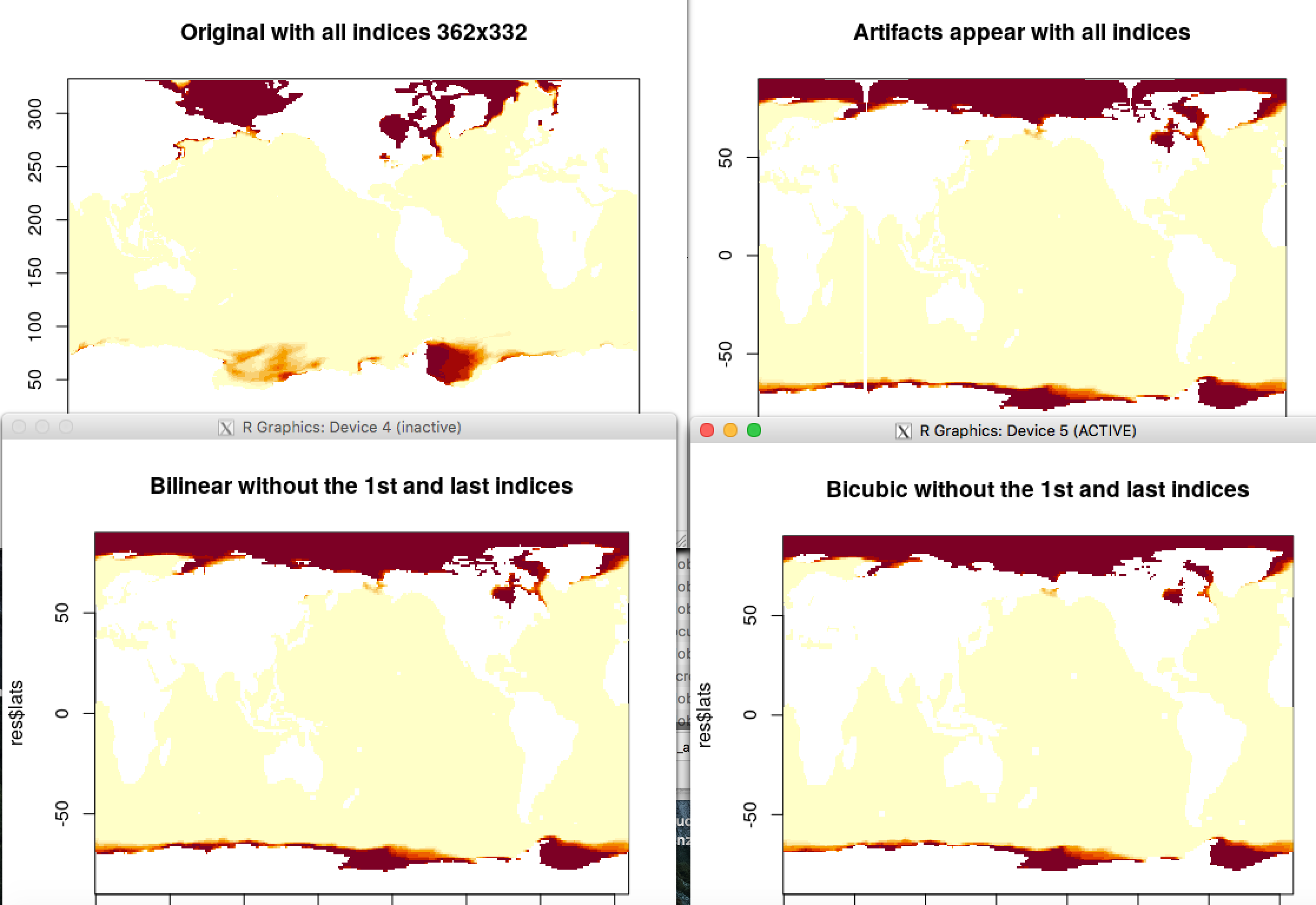
image(res$lons, res$lats, res$data_array, main = "Regrided without the 1st and last indices")The figure shows the original data (top-left), the artifacts (top-right) and the results excluding some indices before the interpolation with 'bicubic' or 'bilinear' methods.
This issue is related to s2dverification#259 (closed) but at that time the files were processed by @pabretonniere and Marga. It may also be interesting for @cdelgado.
Please, Roberto, let me know about the case of CMCC data, so, we can report it here.
With @aho I would like to discuss in a chat the possibility of creating a FAQ and the possibility of using 'transform' parameters in Start, instead of using CDORemap.
Please, don't hesitate to add here any information you may found important.
Thanks, especially to @rfernand for his patient.
Núria